Introduction
Protein domain analysis is a fundamental part of bioinformatics. Domain analysis allows scientists to understand the function of a protein without explicitly testing it in a laboratory setting, or a least get an approximation of the function. When working across taxa with a complex protein structure, domain analysis can also help indicate which protein domains are most crucial as they will likely be the most conserved ones. PFAM is a marvelous tool for doing these analyses, as it combines all of the information into an easy-to-use website with a user-friendly interface [1].
Results and Discussion
|
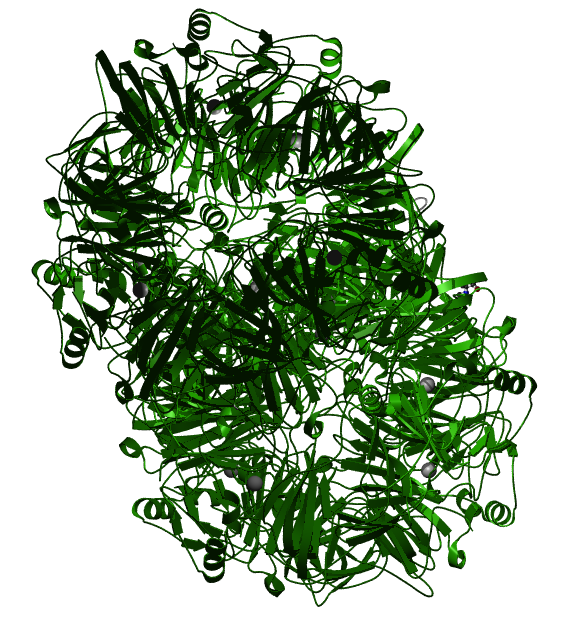
HGD is different from most genes in that it consists of only one domain. This domain spans the entire length of the protein, and is responsible for the main function of the protein: the cleaving of the aromatic ring in tyrosine and phenylalanine [2].
|
Fig 1: The three-dimensional structure of the HgmA domain, which is the only domain found in the HGD protein [2].
|
Conservation of HGD protein structure across taxa
Owing to its importance to general biological function, HGD is the same across most animal taxa, again consisting of only one HgmA domain. While this result means that domain analysis will not be particularly effective for this gene, it does not discount the technique as a whole. Because the HGD protein is so simple and so highly conserved, it means that general amino acid motifs in the sequence can be analyzed to determine the most important regions of the protein. To learn more about this process, please visit the "Phylogeny" page under the "Protein" tab.
Fig 2: A diagram depicting the differences, or in this case, the lack thereof, in the HGD protein structure across popular model organisms. The organisms shown are humans, the house mouse, the brown rat, the fruit fly, the cow, and the horse.
References:
[1] Bateman, A., Coin, L., Durbin, R., Finn, R. D., Hollich, V., Griffiths‐Jones, S., ... & Studholme, D. J. (2004). The Pfam protein families database. Nucleic acids research, 32(suppl_1), D138-D141.
[2] Titus GP, Mueller HA, Burgner J, Rodriguez De Cordoba S, Penalva MA, Timm DE; , Nat Struct Biol 2000;7:542-546.: Crystal structure of human homogentisate dioxygenase.
[1] Bateman, A., Coin, L., Durbin, R., Finn, R. D., Hollich, V., Griffiths‐Jones, S., ... & Studholme, D. J. (2004). The Pfam protein families database. Nucleic acids research, 32(suppl_1), D138-D141.
[2] Titus GP, Mueller HA, Burgner J, Rodriguez De Cordoba S, Penalva MA, Timm DE; , Nat Struct Biol 2000;7:542-546.: Crystal structure of human homogentisate dioxygenase.