Introduction
A protein interaction network is a visualization tools that shows associations between proteins. These associations are often labeled by gene ontology terms as well, so that the relationships are easier to make sense of. Protein interaction networks are great tools for researchers looking for proteins related to their gene/phenotype of interest that might otherwise be unknown. Because of this, studying protein interaction networks can be a good "jumping-off" point when looking for a new avenue for research. Due to the nature of bioinformatics, researchers often uncover associations they would never have time to look into even if they had time to; the fact that protein interaction networks can allow researchers who may be interested in a previously ignored relationship means less of that effort it "wasted".
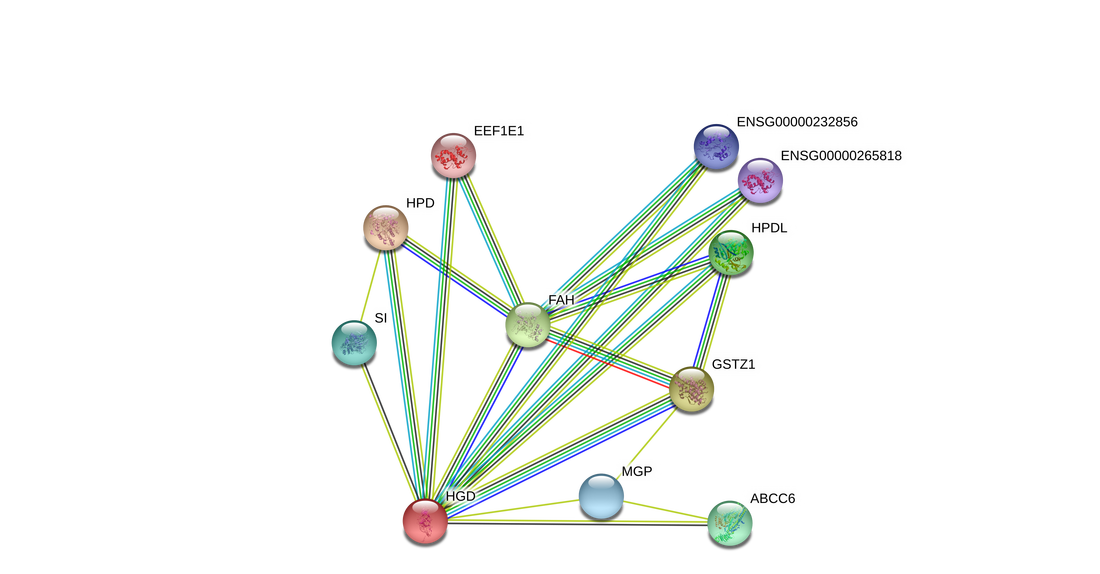
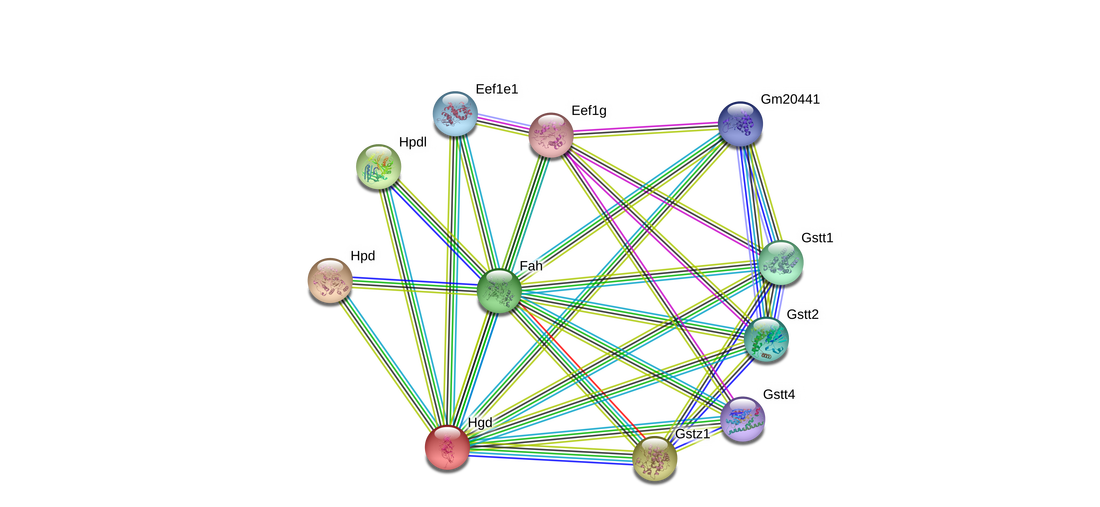
Figure 1: The protein interaction network for HGD (humans) on the left, and HGD (mouse) on the right [1]. There is a large amount of similarity between the two networks, which is reasonable given the highly conserved nature of the protein.
Results and Discussion
|
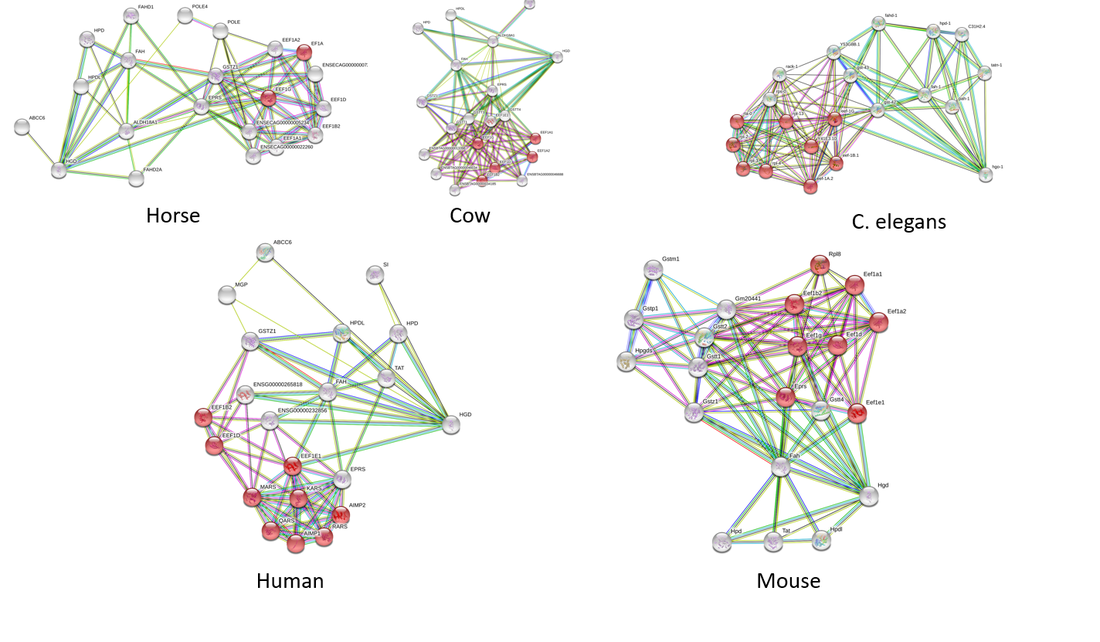
HGD showed somewhat similar protein interaction networks across several model organisms. All showed strong interactions with proteins involved in amino acid metabolism, cell metabolic processes, and other gene ontology terms synonymous with HGD's primary function of tyrosine and phenylalanine catabolism. However, once the network is expanded to see more connections, a strong trend towards translation-related processes appears. In addition, synthesis of tRNA molecules also begins to show up. I cannot think of why translation might be associated with HGD, so I will not be using that connection in my project. However, this connection may still be interesting to any researcher looking to investigate translation changes due to disease states.
|
|
References:
[1] Szklarczyk D, Morris JH, Cook H, Kuhn M, Wyder S, Simonovic M, Santos A, Doncheva NT, Roth A, Bork P, Jensen LJ, von Mering C. The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 2017 Jan; 45:D362-68 |
Image references:
Header: https://bmm.crick.ac.uk/~bmmadmin/Affinity/all_lowdef.jpg |