Introduction
|
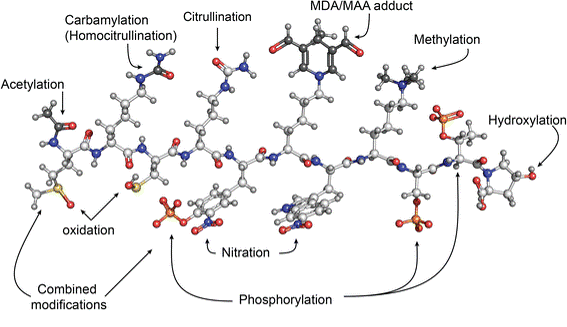
Post-translational modifications is a broad term used to describe the changes seen in a protein between its translation and its functional state. These changes are very diverse, and include a variety of chemical additions, as seen on the right.
Not only are these modifications important for the functioning of the end-product protein, but they can be a useful tool for researchers. Because the modifications can be important for protein function, a change in the modifications could be a contributing factor to a disease state or some other change in the normal functioning of the protein. Analyzing changes in these modifications can also give researchers a better insight into how different mutations affect protein function. Even though a change in a gene may only change one amino acid in a seemingly insignificant part of a protein, that site could be an important phosphorylation site that has now been "deactivated". |
Figure 1: Some of the common post-translational modifications
|
Results and Discussion
|
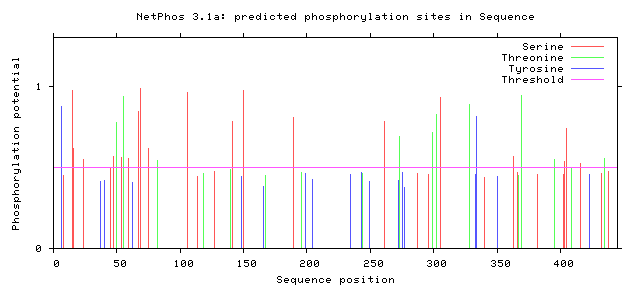
For HGD, I decided to specifically examine phosphorylation, as it is a very common and very important protein modification that was likely going to be found on HGD. I used NetPhos, a tool that uses neural network algorithms to predict phosphorylation sites based on protein sequences [1]. The results of the algorithm showed the potential for a variety of phosphorylation sites on the human HGD gene. This result is exciting, as there are a large number of documented HGD mutations that cause alkaptonuria, some with different severity than others. Part of this could be due to mutations interrupting phosphorylation sites of different levels of importance.
Because HGD is a relatively simple protein, in that it isn't membrane bound, is relatively small, and is made of only one major domain, I did not investigate other post-translational modifications very deeply. |
Figure 2: The NetPhos 3.1 results for the human HGD protein sequence [1].
|
|
References:
[1] Sequence- and structure-based prediction of eukaryotic protein phosphorylation sites. Blom, N., Gammeltoft, S., and Brunak, S. Journal of Molecular Biology: 294(5): 1351-1362, 1999. |
Image references:
Header: http://www.interactive-biology.com/wp-content/uploads/2012/05/Protein-Structure-1280x640.jpg Figure 1: https://media.springernature.com/full/springer-static/image/art%3A10.1186%2Fs12859-014-0370-6/MediaObjects/12859_2014_Article_370_Figa_HTML.gif |